Anna Tovo, Samir Suweis, Marco Formentin, Marco Favretti, Igor Volkov, Jayanth R Banavar, Sandro Azaele, Amos Maritan
Science Advances. Vol. 3, no. 10, e1701438 (2017)
DOI: 10.1126/sciadv.1701438
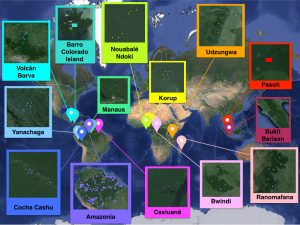
The challenge of estimating global tropical species richness. A map depicting the 15 forests in our data set in which the coordinates of each subplot (squares) are known. Our goal is to deduce the species richness and abundances of each entire forest on the basis of the very limited knowledge in the marked dots (see Table 1 and section S6 for a more detailed description of the data set). From Tovo et al., Science Advance 2017

Leave a Reply
You must be logged in to post a comment.